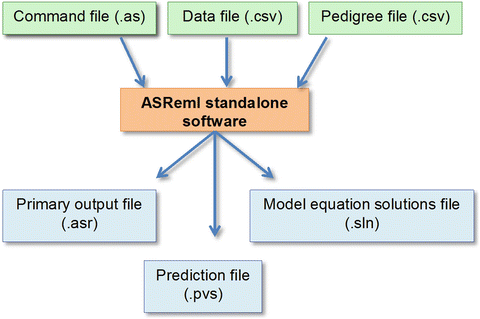
If you think about it, spending a couple of grands in software (say ASReml and Mathematica licenses) doesn’t sound outrageous at all. At that point, of course, I want to be able to get the most of my data, which means that I won’t settle for a half-assed model because the software is not able to fit it.
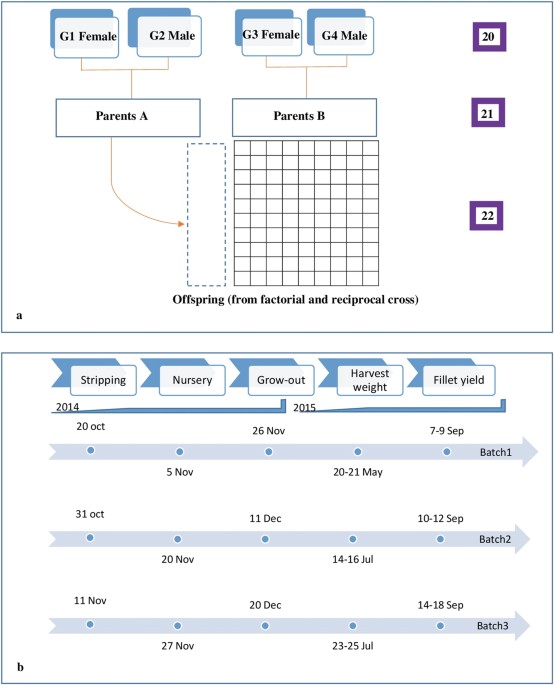
By now you get the idea, actually replicating the research may take you quite a few resources before you even start to play with free software. We took a few hundred increment cores from a sample of trees and processed them using a densitometer, an X-Ray diffractometer and a few other lab toys.
#ASREML UNCONSTRAIN CODE#
Nevertheless, and here is the BUT coming, there is a large difference between making the code repeatable and making research reproducible.Īs an example, currently I am working in a project that relies on two trials, which have taken a decade to grow. There has been enormous progress in the R world on literate programming, where the combination of RStudio + Markdown + knitr has made analyzing data and documenting the process almost enjoyable.

Even here, many people focus not on the models per se, but only on the code for the analysis, which should only use tools that are free of charge. While I share the interest in reproducibility, some times I feel we are obsessing too much on only part of the research process: statistical analysis. In many topics we face multiple, often conflicting claims and as researchers we value the ability to evaluate those claims, including repeating/reproducing research results. There is an increasing interest in the reproducibility of research. This post is tangential to R, although R has a fair share of the issues I mention here, which include research reproducibility, open source, paying for software, multiple languages, salt and pepper. M2 = MCMCglmm(strain ~ species, random = ~ fam + row + column,

